As the literature on exosomes continues to grow in cancer basic science, important questions remain regarding opportunities in the clinic
Researching extracellular vesicles (EVs), including exosomes, can enhance the current biological understanding of the processes involved in cancer development and progression. These small particles, in fact, play a key role in mediating intercellular cross-talk in normal physiology and in pathology. “The interest of researchers is high, and EVs are being studied across the spectrum of cancer, from precancerous states, to localised disease and to the creation of the pro-metastatic niche,” explains Prof. Christian Rolfo, Associate Director for Clinical Research at The Center for Thoracic Oncology, Tisch Cancer Institute at Mount Sinai, New York, USA and President of the International Society of Liquid Biopsy.
“What we know today is, for example, that patients with chronic obstructive pulmonary disease who later develop lung cancer have noticeable alterations in their EVs. At the other end of the scale, in advanced cancer, we know from preclinical models that certain proteins in exosomes can promote brain vascular remodelling and metastasis. Depleting this protein can disrupt invasion and tumour cell association within the brain vasculature, reducing brain metastasis. Findings such as these may help identify new therapeutic targets.”
But do exosomes play a role in the clinic yet? One of the areas of interest is their utility as prognostic markers. As an example, Rolfo describes his recent study that evaluated the predictive role of PD-L1 and TGF-β in EVs from lung cancer patients receiving immune checkpoint inhibitors (J Clin Oncol 2021;39(15 Suppl):e21144). “We found that the kinetics of EV PD-L1 may serve as a better predictive factor than tissue PD-L1. We also reported that increasing dynamics of EV PD-L1 and high baseline EV TGF-β were associated with shorter progression-free survival and overall survival.”
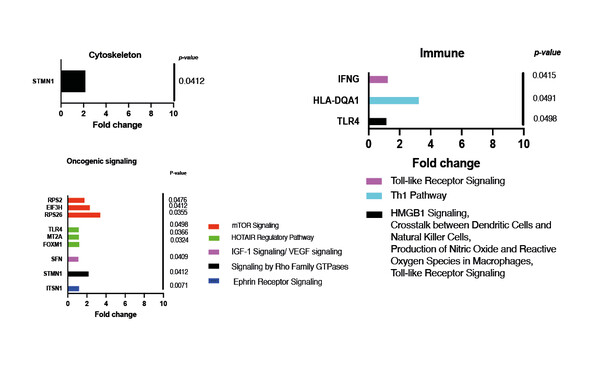
An e-Poster presented at the Molecular Analysis for Precision Oncology Virtual Congress 2021 (MAP 2021 Virtual) used a wide-reaching approach to screen for EV pathway changes predictive of response to cabazitaxel in patients with metastatic castration-resistant prostate cancer (mCRPC) (Abstract 74P). Researchers found significantly upregulated oncogenic signalling, cytoskeleton- and immune-related pathways at baseline in non-responders. In contrast, responders exhibited differential alterations including upregulated NK signalling. The levels of enriched pathways in non-responders at baseline did not show any significant change after the first cycle, whereas there were significant changes over time in responding patients.
“Using two different platforms for interpreting and capturing information at different time points are important strengths of this research, although the number of patients was small,” explains Rolfo. “I do not believe that information about EVs will completely replace as a biomarker other analyses of the liquid biopsy family, such as ctDNA that provides robust data on a cancer, but I think that EVs can add to this biological information in a clinically meaningful way.”
We are currently missing patients with minimal residual disease using existing methods. Integrating exosomes into a multi-omics approach could therefore be an important future application.
Also presented at MAP 2021 Virtual is the design of an ongoing study that is currently investigating the use of exosomes in monitoring response to neoadjuvant chemotherapy of patients with pancreatic ductal adenocarcinoma (Abstract 84TiP). Changes in exosome number, size and cargo will be compared prior to and during neoadjuvant chemotherapy in patients who recurred versus those who remained with no evidence of disease during follow-up.
Rolfo thinks this study will add to existing research in early disease in other tumour types. “We have previously shown that liquid biopsies based on exosomal micro (mi)RNAs and circulating tumour cells can be complementary clinical tools in the diagnosis and prediction of treatment response in localised breast cancer undergoing neoadjuvant chemotherapy (Breast Cancer Res 2019;21:21),” he says. “We are currently missing patients with minimal residual disease using existing methods. Integrating exosomes into a multi-omics approach could therefore be an important future application.” In addition, Rolfo highlights the part that exosomes may play in mediating resistance to therapies, the monitoring of which may be an important early step to guide subsequent treatment choices.
Technical aspects, such as lack of automation for purification and isolation, and inadequate standardisation, are currently limiting wider research and use of exosomes, according to Rolfo. With further development, however, he thinks that there are other opportunities for exosome research beyond biomarkers. “Due to their role as important intracellular communicators, exosomes are natural vehicles – they could be labelled for imaging or engineered for delivery of anticancer drugs or siRNAs. Using exosomes to re-direct cells away from tumourigenesis or metastasis could be a powerful therapeutic option for the future.”
Abstracts presented:
Vardaki I et al. Exosome profiling reveals prognostic markers of response to Cabazitaxel in metastatic castration resistant prostate cancer (mCRPC). MAP 2021 Virtual, Abstract 74P
Cazacu IM et al. Predictive Value of Exosomes for Therapy Response in Resectable/Borderline Resectable Pancreatic Cancer Patients. MAP 2021 Virtual, Abstract 84TiP