Three studies support the use of NGS to better understand the molecular drivers of this brain tumour, but effective targeted treatments are still lacking
Results from several studies presented at ESMO Congress 2022 indicate that molecular alterations can be identified in glioma by next-generation sequencing (NGS), paving the way for the development of effective targeted therapies for these tumours
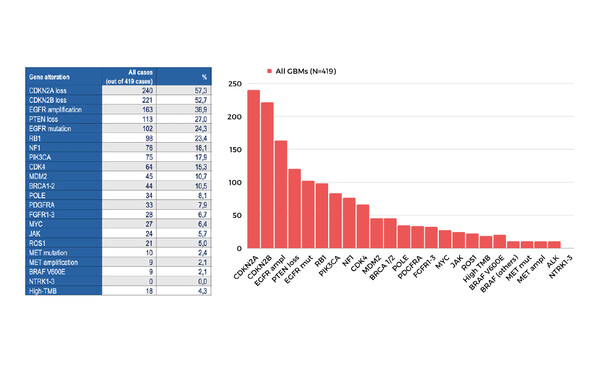
At the Congress, Padovan and colleagues presented the results of two single-centre studies on the use of NGS in glioma. In a Mini Oral Session (Abstract 289MO), they described how NGS is a feasible tool to evaluate glioblastomas and may identify a large number of patients who are eligible for personalised treatment either with licensed or investigational therapies. In this study, an NGS profile was obtained for 351 primary and 68 recurrent tumour samples from patients with glioblastomas that were wild-type for isocitrate dehydrogenase (IDH). Actionable molecular alterations were frequently observed, including CDKN2A loss (57% of patients), CDKN2B loss (53%), EGFR amplification/mutation (39/24%) and PTEN loss (27%), although no NTRK1/2/3 druggable alterations were detected. Statistically significant differences in the incidence of EGFR, RB1 and PIK3CA alterations were also observed between the subgroups of primary and recurrent tumours. The research team also presented a poster (Abstract 297P), which reports that NGS can provide evidence of genetic profile changes during glioma progression. This study evaluated 46 matched primary and recurrent glioma samples from 23 patients, all of whom had received temozolomide with or without radiotherapy after their initial surgery. De novo gene alterations, including mutations in TP53, loss of PTEN and alterations in NF1, were detected in the recurrent tumours in seven patients. In a further six patients, genetic alterations detected in the primary samples (CDKN2A loss [n=2] and EGFR alterations [n=4]) were no longer observed after recurrence. Median tumour mutational burden was considerably high in the recurrent versus primary samples (14.9 versus 1.68 mutations/megabase), consistent with treatment-induced hypermutagenesis.
Also presented as a poster at ESMO Congress 2022 were the results from a single-centre study demonstrating a high prevalence of ESMO Scale for Clinical Actionability of molecular Targets (ESCAT) tier I–II alterations in primary brain cancers (pBC), indicating that molecular profiling should be offered to these patients (Abstract 290P). In this study, NGS was performed for 159 patients with a pBC. Overall, 15.1% of patients had a tier I or II alteration, with tier I defined as alterations suitable for routine use of targeted treatment and tier II defined as alterations that are associated with treatments having antitumour activity, but where the magnitude of clinical benefit is unknown. This included four patients with a tier I alteration (BRAF V600E mutation) and 25 with a tier II alteration (PTN loss [n=3], IDH1 mutation [n=16]. A further 48 and 40 patients had tier III or IV alterations, respectively. This study also identified potential enrichment factors, such as non-glioblastoma histology, age (<40 years) and IDH1 mutation subtype, that may be used to optimise which patients should be offered NGS.
According to Prof. Martin van der Bent from the Erasmus Medical Centre, Rotterdam, Netherlands, NGS is now frequently used as a standard diagnostic tool in many centres as it allows common tumour mutations to be identified with high accuracy and specificity in a single analysis. “However, the clinical utility of this technology in the treatment of glioma remains unanswered at the current time,” he adds. “Many molecular targets in glioma can be identified but, with the exception of BRAF, these do not currently lead to meaningful clinical treatments.” For van der Bent, this is partly due to the fact that in some targets, genetic alterations and their clinical significance can vary between tumour types. “For example, although EGFR mutations occur commonly in glioma, established EGFR inhibitors such as osimertinib and erlotinib are generally not effective as the specific mutations they target are rarely found in this setting,” he says. “Furthermore, even if effective agents are developed for specific molecular targets, the benefits will need to be elucidated for specific glioma patient subpopulations. It will be essential to identify targets and then treat glioma patients in the context of a clinical trial or registry in order to fully understand how the use of targeted therapies can be optimised. It is simply not enough to identify a target; they must lead to clinically meaningful treatments.”
Abstracts presented:
Padovan M, et al. Next-generation sequencing (NGS) for identifying actionable molecular alterations in newly diagnosed and recurrent IDHwt-glioblastoma (GBM) patients: A large mono institutional experience. ESMO Congress 2022, Abstract 289MO
Mini Oral Session 10.09.2022, h. 16:30 – 18:00, Urval Auditorium
Padovan M, et al. The genetic profile of primary and recurrent gliomas: A mono institutional experience using next-generation sequencing. ESMO Congress 2022, Abstract 297P
Poster display, Hall 4. An e-Poster is also available on the Congress virtual platform
Mirallas O, et al. Potential enrichment strategies for next-generation sequencing (NGS) in primary brain cancers (pBCs) in a clinical series according to ESMO scale for clinical actionability of molecular targets (ESCAT). ESMO Congress 2022, Abstract 290P
Poster display, Hall 4. An e-Poster is also available on the Congress virtual platform