Data from two novel studies further support the feasibility of this approach, although its clinical utility has yet to be determined
In early breast cancer, trials testing circulating tumour DNA (ctDNA) in the surveillance setting provide key learnings to further investigate whether this approach is helpful for patients: there is a need for ultrasensitive tests as ctDNA levels can be extremely low and difficult to detect in early-stage disease; and screening should be started early, especially in patients with triple-negative breast cancer (TNBC) where high rates of metastatic disease are found at the time of first ctDNA detection. These are the take-home messages from two oral presentations at the ESMO Breast Cancer 2026 congress (Berlin, 6–8 May), which also highlighted the challenges of conducting clinical trials in this emerging area of research.
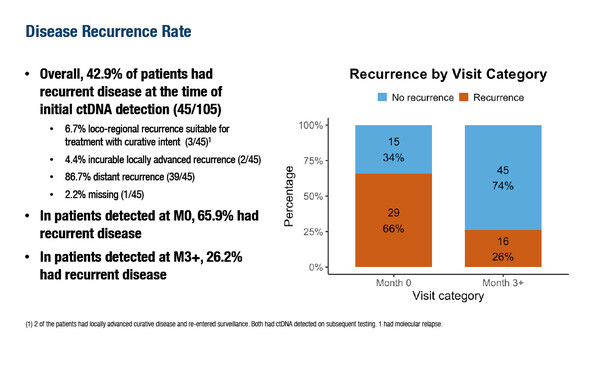
Novel, ultrasensitive tumour-informed assays could enable the early detection of minimal residual disease (MRD), maximise lead time and reduce the rate of recurrent disease at the time of ctDNA detection. As presented at the congress, in the TRAK-ER randomised, multicentre study, ctDNA surveillance every 3 months in patients with high-risk, hormone receptor (HR)-positive, HER2-negative breast cancer receiving adjuvant endocrine therapy showed an overall ctDNA detection rate of 1.87% (107/5,724 samples) (Abstract 1O). The first surveillance test (Month 0) was associated with the highest detection rate (4.96% of samples), and subsequent tests (Month 3+) with a lower detection rate (1.30% of samples). Notably, 42.9% (45/105) of patients overall had recurrent disease at the time of initial ctDNA detection, and of patients with ctDNA detected at Month 0, 65.9% had recurrent disease, decreasing to 26.2% of those with ctDNA at Month 3+.
“The prevalence of ctDNA is very low in early-stage breast cancer, so the number needed to screen to identify positive cases is high. Of the 901 patients with successful panel design and who started ctDNA surveillance in the TRAK-ER study, only 506 (56.2%) had true molecular relapse without clinical or radiographic evidence of disease,” notes Dr Stefania Morganti from the Dana-Farber Cancer Institute/Harvard Medical School, Boston, MA, USA, commenting on the results. “Although ultrasensitive assays are essential to maximise early detection of molecular relapse, the need for tissue represents another limitation of this approach. In TRAK-ER, the assay design failed in 16.1% of patients, highlighting the need for more sensitive tumour agnostic assays that could potentially overcome the limitations of tumour informed tests.” The challenges of utilising ctDNA in the surveillance setting have been reported previously. In the phase II LEADER study, the cumulative ctDNA detection rate during surveillance for patients with stage I–III HR-positive/HER2-negative breast cancer was 11% (15/140) and 33% of the ctDNA-positive cases had overt metastatic disease at the time of ctDNA detection, further underlining the need for ultrasensitive assays in this setting (Clin Cancer Res, 2026;32(4_Suppl): PD5-01). Similar findings were observed in the DARE study, in which 12% of patients were found to be ctDNA-positive during surveillance and 27% of them had overt metastatic disease (J Clin Oncol. 2025;43(16_Suppl):1010). In the phase III ZEST trial of patients with triple-negative breast cancer (TNBC) or HER2-negative, BRCA-mutated breast cancer (Clin Cancer Res. 2025;31(12_Suppl):GS3-01), most patients with a ctDNA-positive test (66%) were positive at the first testing timepoint, suggesting the need to start screening early.
The c-TRAK-TN trial in patients with early-stage TNBC also demonstrates the importance of starting ctDNA screening early and with more sensitive assays; a high rate of metastatic disease (72% of ctDNA-positive patients allocated to the intervention arm) was found at the time of ctDNA detection by digital PCR. As a result, only 5 patients initiated pembrolizumab immunotherapy at molecular relapse and none achieved sustained ctDNA clearance (Ann Oncol. 2023;34:200–211). Further analysis of the c-TRAK-TN study comparing ctDNA detection using more sensitive assays was discussed at the Congress, revealing high concordance (95.2%; 770/809 samples) between a tissue-agnostic approach (based on epigenomic profiling without the need for prior sequencing of the primary tumour) and a multivariant tumour-informed assay (Abstract 4RO). At the patient level, only 7.5% (10/133) had discordant results. Morganti notes that of the ctDNA-positive cases detected by both assays, almost one-third (29.3%) were detected at an earlier timepoint by the tumour-informed assay, though the median lead time (from first ctDNA detection to recurrence) was similar between the two approaches: 7.1 months with the tumour-informed assay and 7.6 months with the tissue-free assay (hazard ratio 1.46; 95% confidence interval 0.87–2.44; p=0.154).
“Tumour-informed assays with greater sensitivity in detecting ctDNA are now available and it is unknown how these would compare with the tissue-free assay,” Morganti points out. “In addition, this study was conducted in patients with TNBC, a high-shedding tumour resulting in a relatively high tumour fraction. We do not know if tissue-free assays would be sufficiently sensitive to detect ctDNA in lower shedding tumour types such as HR-positive early breast cancer.”
Multiple studies have shown that detection of ctDNA in patients with early breast cancer is associated with a high risk of recurrence (ESMO Open. 2024;9:102390). However, the clinical utility of this approach has not been demonstrated.
“Before implementing these tests in routine surveillance, we need evidence that we can improve patient outcomes through ctDNA-guided treatment interventions,” concludes Morganti. “Detection of molecular relapse during surveillance anticipates distant recurrence, but research is needed to understand whether MRD is curable in this setting. Without studies proving that intervening at the time of molecular relapse significantly delays or prevents recurrences, these tests should not be used in clinical practice given the absence of proven benefit to outweigh the risk of toxicity related to intervention, the anxiety associated with test results, and their costs.”
At a glance
Cunningham N, et al. ctDNA detection rates during surveillance in high-risk HR+/HER2 negative breast cancer from the TRAK-ER study. ESMO Breast Cancer 2026 - Abstract 1O
- N=901 with ctDNA result available from ≥1 timepoint analysis
- Panel design failure rate: 16.1% (178/1,104)
- ctDNA detection: 1.87% of samples (107/5,724); 11.6% of patients (105/901)
- 42.9% of patients had recurrent disease at time of initial ctDNA detection
- Median allele fraction:
- M0 samples: 0.056%
- M3+ samples: 0.0135%; p<0.001
- 56.2% of patients positive for ctDNA had molecular relapse but no clinical/radiographic evidence of disease
Cunningham N, et al. A comparative analysis between tissue-free ctDNA detection and a multivariant tumour-informed assay in early triple negative breast cancer. ESMO Breast Cancer 2026 - Abstract 4RO
- N=133
- Concordance between tissue-free and tumour-informed ctDNA detection: 95.2% (770/809 samples)
- Discordant results between assays: 7.5% of patients (10/133)
- ctDNA detection by both assays: 30.8% of patients (41/133)
- Earlier detection by:
- Tumour-informed assay: 29.3% of patients (12/41)
- Tissue-free assay: 2.4% (1/41)
- Earlier detection by:
- Median time from first ctDNA detection to recurrence:
- Tumour-informed assay: 7.1 months
- Tissue-free assay: 7.6 months